Apart from finding life, a new ecology study on New York City’s East River also reveals the city’s dining habits. Researchers looked at what’s living in the water over a year and analyzed the DNA from what they found, in living organisms and in remains from human feces. The samples included DNA from chicken, pork and rats.
The researchers took liter water samples every time and revealed in unexpected detail a snapshot of life in and around the city, from weekly and seasonal shifts in fish species and abundance to urban wildlife activity, the pets they keep and what New Yorkers eat.
The findings suggest that urban waterways anywhere could become continuous biosensors, tracking biodiversity, habitat restoration outcomes, and human impacts in real time.
Among the most novel results of the weekly East River sampling were genetic indicators of human food consumption, of rat and other terrestrial wildlife populations, and the discovery of newly abundant fish species, thought to reflect the success of habitat restoration efforts.
“The water that flows under the Brooklyn Bridge and alongside the United Nations Headquarters is teeming with life and information valuable for improving nature where most people live,” said lead author Mark Y. Stoeckle, M.D., of The Rockefeller University’s Program for the Human Environment.
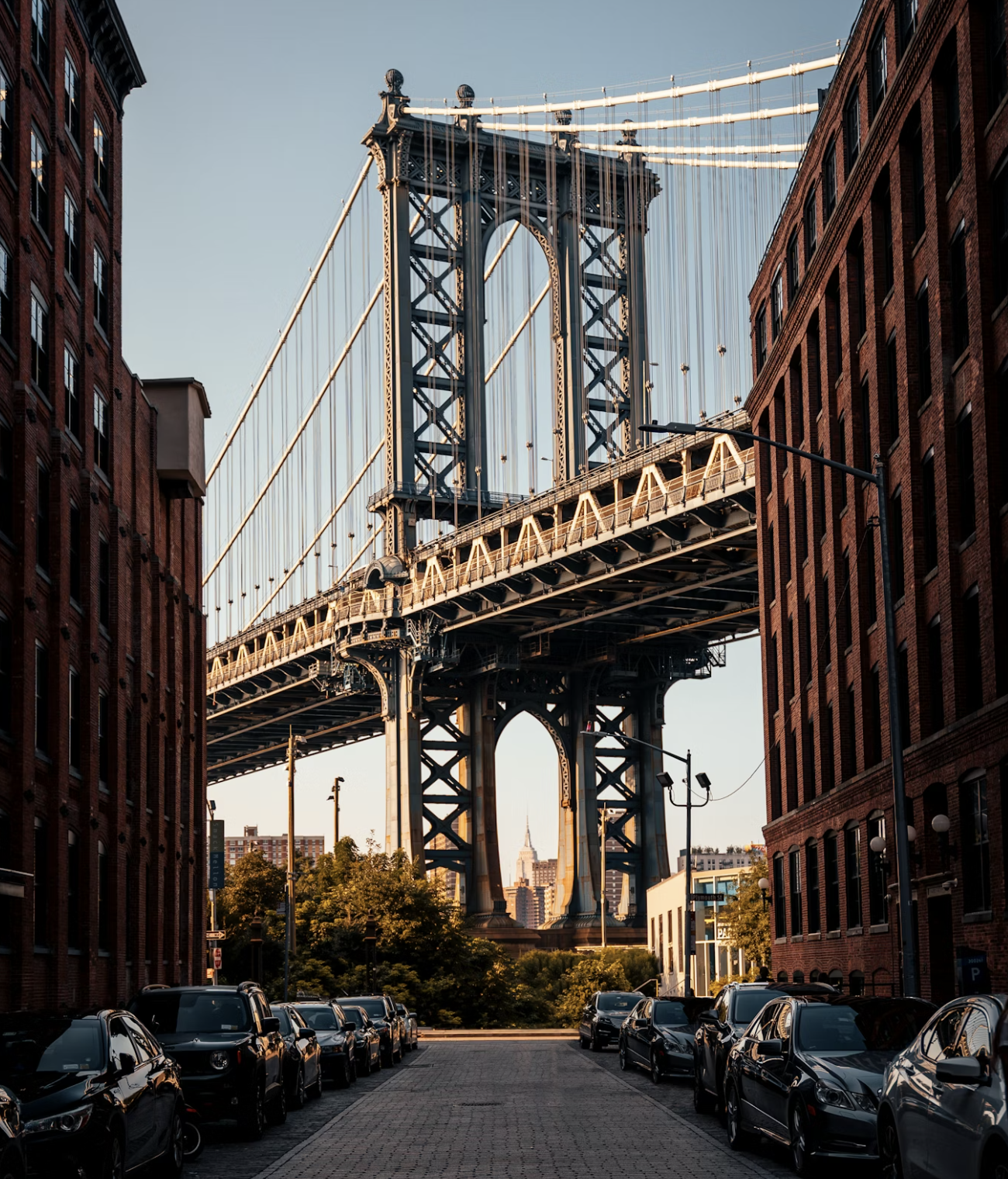

“We can now assay life in and around rivers by applying modern genomic techniques to a small sample of water,” said Dr. Stoeckle. “This approach promises to make environmental monitoring as simple and routine as a blood test.”
“Environmental DNA doesn’t just tell us what lives in the water, it reveals insights into the entire ecosystem surrounding it, including the city itself.”
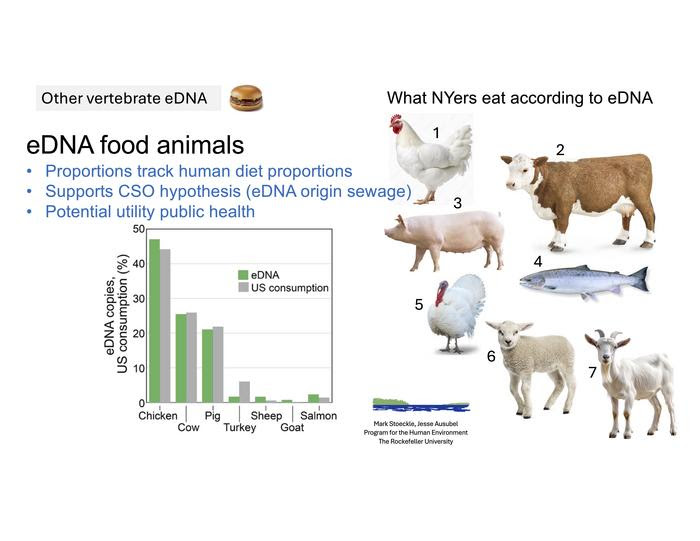
The most unexpected findings, Dr. Stoeckle said, included the detection of food animals’ DNA in proportions closely aligned with human diet patterns – genetic traces of chicken, beef, pork, turkey, lamb, goat, and commonly consumed fish such as salmon and European sea bass.
Instead of obscuring ecological data, these readings provide a unique window into eating patterns and public health trends, said Dr Stoeckle.
From sampling fish to reading the city
About eDNA: Fish and wildlife leave environmental DNA traces in water by shedding skin cells, waste, or other biological traces. Thanks to advances in sequencing technology, researchers can now use these waterborne DNA traces to identify species and estimate their relative abundance, among other insights.
Sampling was simple: weekly 1-liter shoreline water grabs from May 2024 to May 2025 put through something akin to a small coffee filter. From the sediment on a piece of the filter the size of a small tortilla chip, using 12S metabarcoding primers, the researchers produced a continuous, high-resolution picture of seasonal fish abundance.
Urban waterfronts are classically tough places to study for ecological and social reasons, but eDNA makes them safely accessible,” Ausubel added. “With the right coordination, this approach could become the backbone of urban coastal monitoring across the country.
Key patterns:
- Seasonal cycles were clear, with species arriving and departing in predictable waves
- Results aligned with traditional trawl surveys, confirming the method’s reliability
- By monitoring eDNA-dominant species, researchers can identify habitats and flag shifts quickly
“We can turn raw DNA reads into reliable abundance estimates,” Dr. Stoeckle noted. “That brings eDNA out of the experimental phase and into everyday scientific use.”
The East River’s most reliably detected species included skilletfish, tautog, and feather blenny.

Indeed, the study found skilletfish and feather blenny surprisingly abundant compared with earlier monitoring efforts, thought to reflect recent remediation initiatives such as oyster reef restoration.
In all, the eDNA of 71 local marine fish species was detected and changes in their abundance was tracked over time. Fish DNA levels rose roughly tenfold during summer months, consistent with known ecological patterns, demonstrating that eDNA can reliably and accurately reflect biological dynamics.
Many dominant species were reef- or structure-associated fish aligned with the rocky habitat of the East River, reinforcing confidence in the reliability of eDNA signals.
Frequent eDNA sampling could detect ecological change earlier than traditional monitoring programs, offering a near-real-time view of ecosystem response.
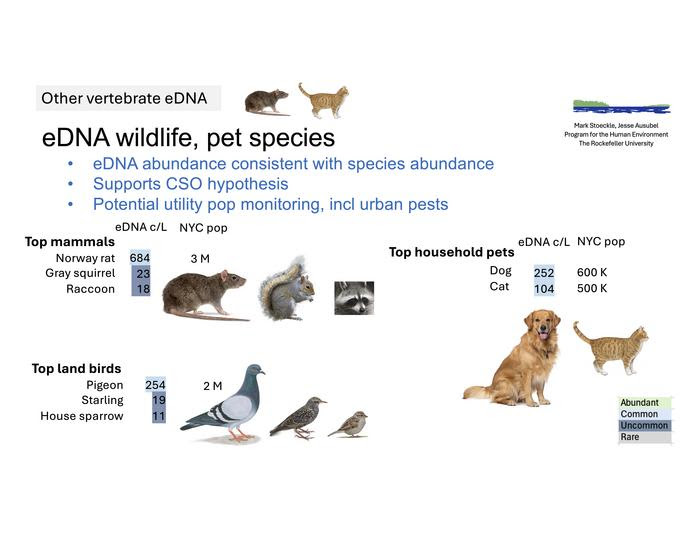
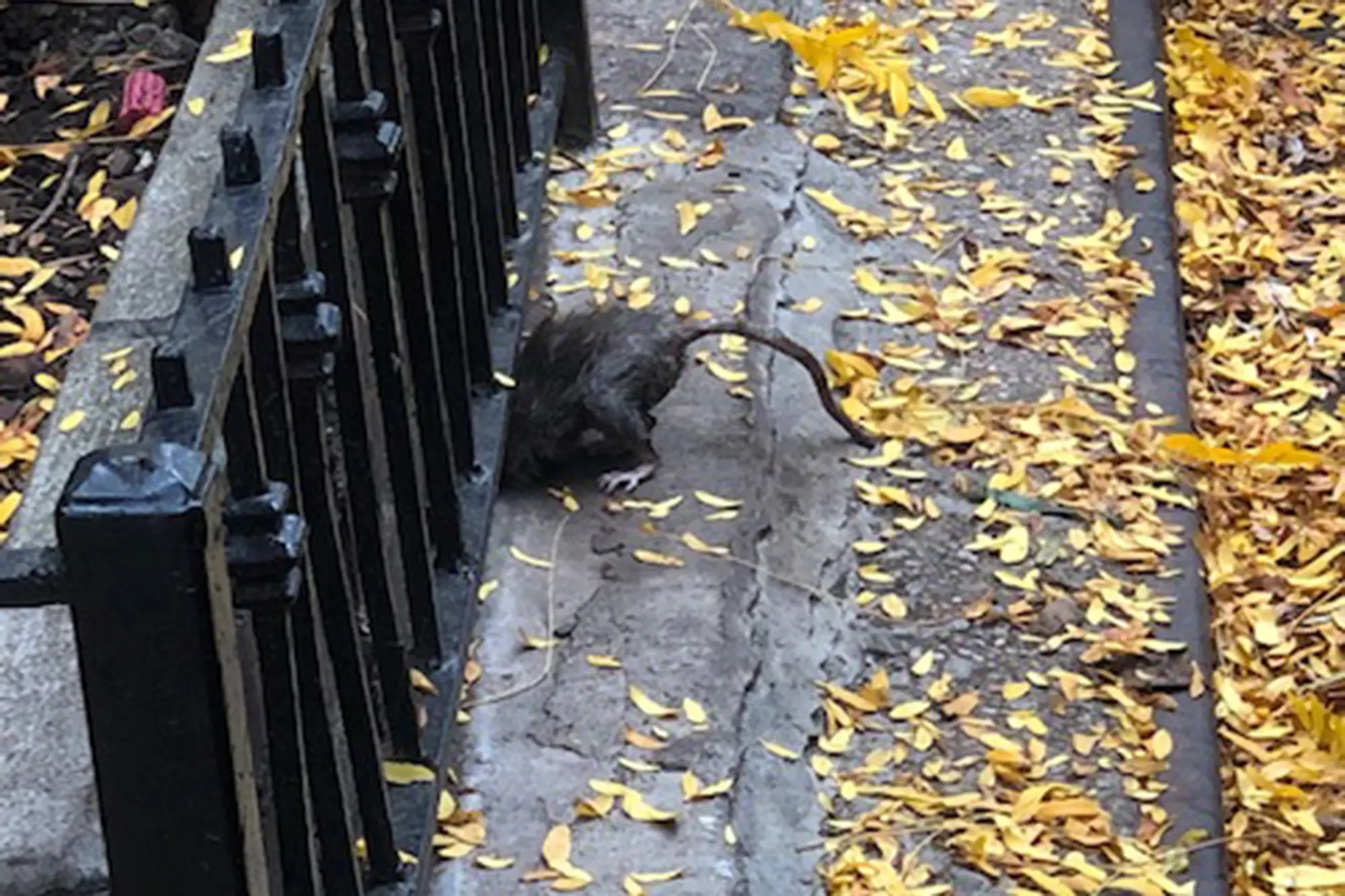
Alongside marine species, the study detected DNA from more than 60 terrestrial animals, including rats, raccoons, squirrels, and common city birds. Levels of wildlife DNA tracked closely with human DNA signals, suggesting combined sewer overflows were transporting biological information into the river.
Using eDNA, municipal authorities could potentially assess, for example, whether rat populations are increasing or decreasing, that is, whether rat control programs are working, Dr. Stoeckle said.
Towards continuous urban environmental monitoring
“After a heavy rain, the DNA of almost everything that makes the city tick — and squawk and squeak — ends up in the East River,” said co-author Jesse Ausubel, Director of The Rockefeller University’s Programme for the Human Environment.
“Genetically speaking, a rainstorm turns the river into something akin to Times Square on New Year’s Eve: crowded, noisy, and full of signals.”
“Environmental DNA could form the basis of future monitoring, allowing cities to track ecosystem health continuously without traditional surveys that are often limited by cost, logistics, or safety.”
“Urban waterways worldwide could become distributed observatories of ecological change, reporting almost real-time what lives in and near them, not only fish but bats, beavers, and foxes.”
“Urban biodiversity monitoring could expand dramatically and inexpensively using minimal equipment at relatively low cost,” added Dr. Stoeckle. “This ability to integrate environmental and human signals positions eDNA as a powerful tool for understanding the Anthropocene – the era defined by human influence on Earth’s systems.”
Compared with trawl surveys, eDNA detected more species and captured short-lived seasonal pulses that nets simply cannot. Traditional gear remains essential for age structure, reproduction, contaminants, and health, but eDNA reduces “false absences” and fills important gaps.
A full year of weekly sampling cost about US$ 15,000 and a fraction of a staff member’s time – far below the cost of large survey vessels. The method is equally suited to surf zones, shallow estuaries, and other places where nets cannot go. A person, or even a drone, with a bucket on a rope can collect the water sample.
“Urban waterfronts are classically tough places to study for ecological and social reasons, but eDNA makes them safely accessible,” Ausubel added. “With the right coordination, this approach could become the backbone of urban coastal monitoring across the country.”